The following is reprinted from an interview produced by Nature Portfolio on a project in the David Lab sponsored by the Global Grants for Gut Health.
Lawrence David, Sylvia Becker-Dreps, Teresa McDonald and colleagues, together with Filemon Bucardo and Samuel Vilchez in Nicaragua, will use their Global Grant for Gut Health to measure the diversity of plant-based foods consumed by infants in Nicaragua, and how this modulates the maturation of the gut microbiome and overall child health.

Why are plants important to the infant gut?
In adults, a diet with a greater diversity of plants has been associated with a more diverse microbiota, which in turn is often associated with good health. In infants, diverse diets are associated with a lower risk of developing allergies later in life. A main component in plant-based foods is dietary fibre (complex carbohydrates), but humans lack the enzymes to break fibre down — we can’t absorb it like we do simple sugars or proteins. Plant fibre survives ingestion and travels to the large intestine, where most gut bacteria reside. Fibre is an important resource for these bacteria; when deprived of fibre, they eat more intestinal mucus. This can compromise the function of the intestinal barrier and increase the risk of pathogens taking hold. But we’re still learning exactly how these dietary components shape the development of the infant and child microbiome, and whether specific plants benefit young children more than others. As it is feasible to rapidly change the composition of the microbiome in a few days, interventions could work very quickly to improve infant gut health.
Please outline your proposed project.
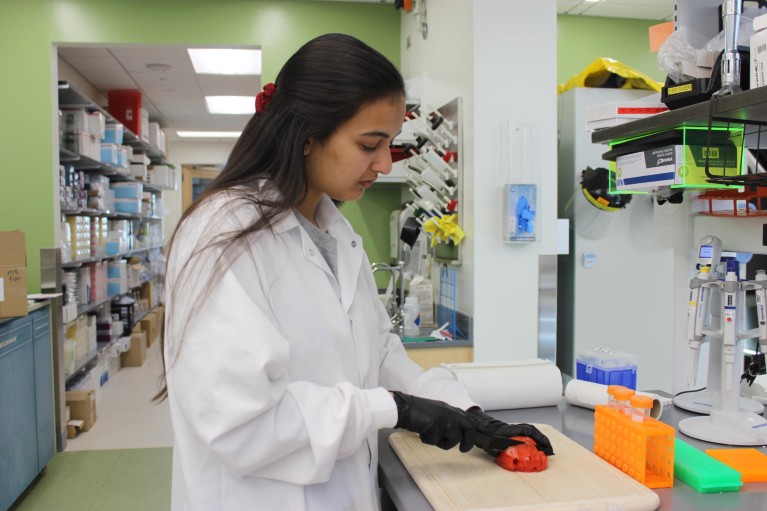
Childhood and infancy are critical periods during the human lifespan, yet we know very little about gut microbiome development. Our project will focus on children from a community in Nicaragua who are at risk of diarrheal illness. Sylvia Becker-Dreps, the physician scientist shaping the clinical aspects of our work, together with Nicaraguan virologist Filemon Bucardo and bacteriologist Samuel Vilchez, has spent nearly two decades examining factors that affect infectious disease risk among kids in Nicaragua and other countries. In our proposed project here, we will measure the diversity of foods that the infants eat as they grow, and connect this to the maturation and composition of the gut microbiome and overall infant health. Children can’t record what they eat, and they’re often fed by multiple caregivers during any given day. This is where our new metabarcoding tool comes in, which uses DNA sequences of stool samples to reconstruct dietary intake without needing participants to record what they eat. We will also test some of the hypotheses generated from these data by growing bacteria on nutrients extracted from food in a human-gut model.
How does metabarcoding work?
The technique was inspired through collaboration with ecologists who have been studying the diets of large herbivores in the African savannah. They couldn’t ask animals what they were eating, so instead they sequenced the residual DNA originating from the animals’ diet that are left behind in their stools. The DNA ‘tags’ in stools can be matched to a library of different foods — both vegetation and meats — allowing the scientists to infer the animals’ diet. We’ve adapted this process for people. We sequence DNA from the chloroplasts of plants and the mitochondria of animals found in human stool samples. Once we have the DNA sequences, we compare them to a large database of different food groups — at present, our library consists of more than 400 common plants and animals. From this, we can infer what young children have been eating.
What challenges are inherent in this project?
There are thousands of edible plant species across the world, so there is still much work to do to gather all the DNA sequences we might find in different diets in different cultures. In our recent pilot study for this project, one sequence found around 20% of stool samples didn’t match any in our databases. One of our lead scientists, Teresa McDonald, did some critical detective work. The fragment was so short that it mapped to multiple plant species, but the only plant species that were both theoretically edible and found in Nicaragua were duckweed and a tuber called malanga. Teresa asked her colleagues at the National Autonomous University of Nicaragua in Leon, and found out that malanga is one of the region’s most popular baby foods. The tuber is grown locally year-round and is believed by residents to be beneficial for infant digestion. This process, in which members of our team from different parts of the world learn about each other’s dietary practices, has been an important and enjoyable process. We’re also still learning how different food processing methods — foods can be eaten raw, boiled, baked or fried — affect our metabarcoding technology. This could influence our ability to infer an overall nutritional picture from DNA.

What will human-gut modelling contribute?
Our artificial gut model is based on pioneering work done by other microbiome groups over the past decades. We place faecal bacteria as a culture media in a vessel, add individual dietary nutrients to the bacteria and then watch what happens. We can mimic ingestion, digestion and defecation with the model. This has several advantages. Firstly, we can fully control the environment, recreating certain parameters of the gut ecosystem and manipulating them in ways that are impossible in a mouse or human subject. We can also study the dynamics of microbial communities over time, and examine the responses of bacteria to different stimuli over minutes or hours.
What practical applications will this project have?
We hope to provide scientific evidence to inform dietary guidelines for children in low and middle-income countries. We recently shared some preliminary data with our collaborators — Vilchez said he had never seen data that characterized Nicaraguan diets in such a granular fashion. If we show these data to parents and connect it to disease risks, they would likely be receptive to it, because the data are from their own community, are easily understandable, and relate to what they care about. It’s also valuable to confirm which foods are incorporated into children’s diets in specific places — this can feed directly into community health and education projects.
What plans do you have for future research?
Even if we pinpoint ways to improve the health of children’s microbiomes, there are still many challenges with changing diet in desired ways. Socio-economic challenges, psychological and behavioural changes — everyone wants to eat well, but it’s often easier said than done. If we can develop ways to help people know that the foods they eat benefit their bodies — by say monitoring the microbiome say on a regular basis — this would be a significant breakthrough.